Cutadapt remoYes adapter sequences from high-throughput sequencing reads Introduction Implementation Features
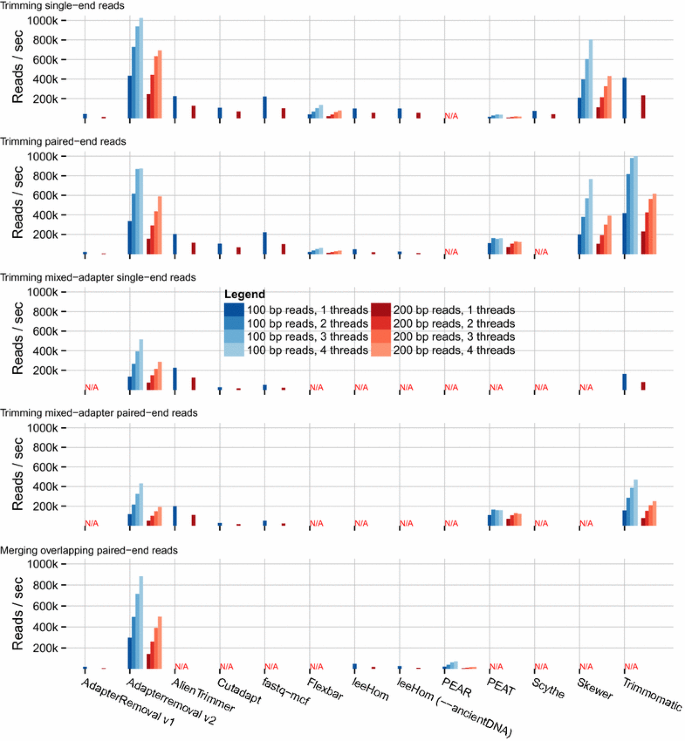
AdapterRemoval v2: rapid adapter trimming, identification, and read merging | BMC Research Notes | Full Text

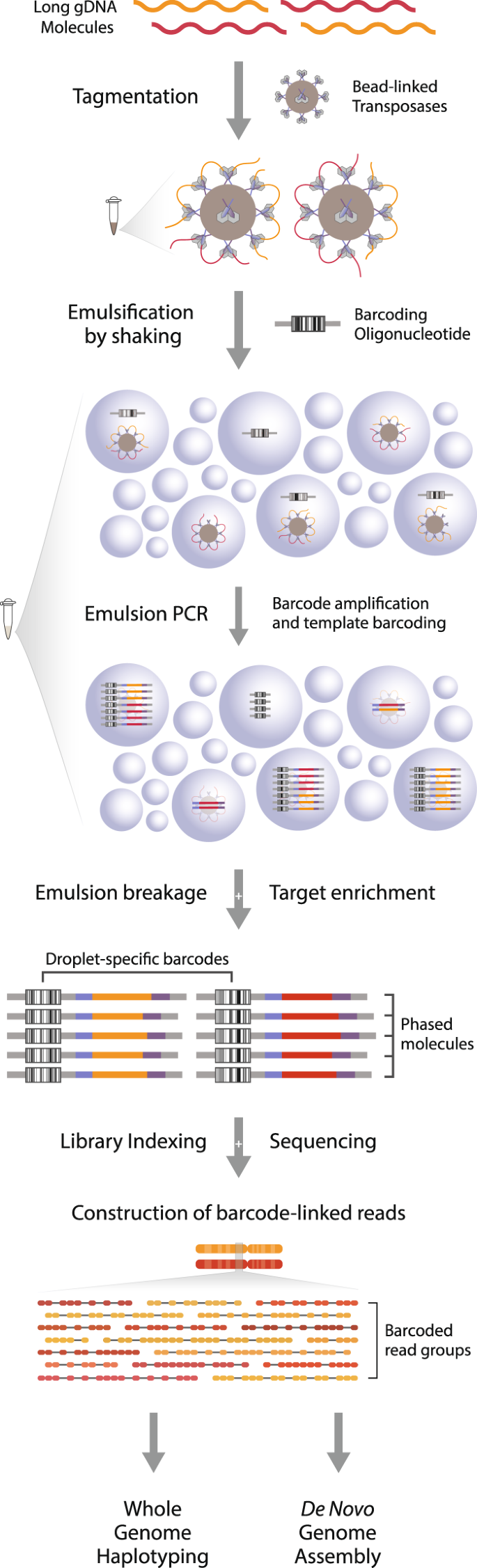
Assessing the utility of long-read nanopore sequencing for rapid and efficient characterization of mobile element insertions - Laboratory Investigation
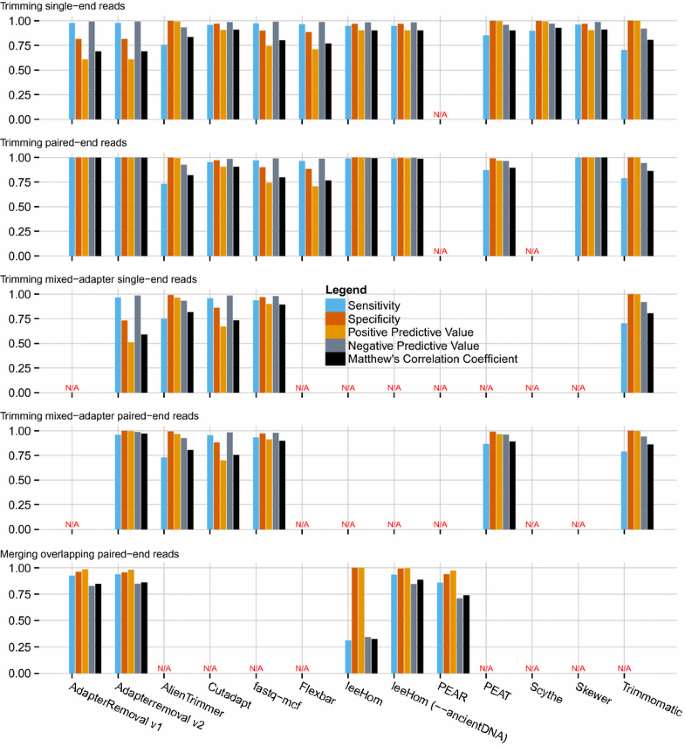
AdapterRemoval v2: rapid adapter trimming, identification, and read merging | BMC Research Notes | Full Text